Machine Learning - Linear Discriminant Analysis
This method is only available in Q5.
Fits linear discriminant analysis (LDA) to predict a categorical variable by two or more numeric variables. Ordered categorical predictors are coerced to numeric values. Un-ordered categorical predictors are converted to binary dummy variables.
The parameters of the discriminant functions can be extracted with Machine Learning - Diagnostic - Table of Discriminant Function Coefficients.
Usage
To run a Linear Discriminant Analysis:
1. In Displayr, select Insert > More > Machine Learning > Linear Discriminant Analysis. In Q, select Create > Classifier > Linear Discriminant Analysis.
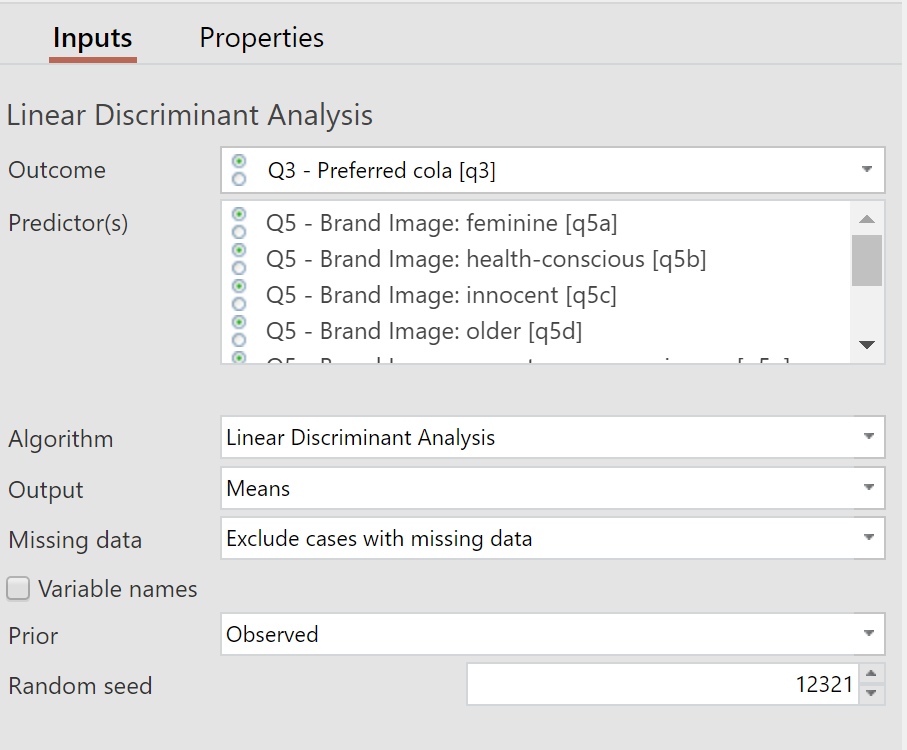
2. Under Inputs > Linear Discriminant Analysis > Outcome select your outcome variable.
3. Under Inputs > Linear Discriminant Analysis > Predictor(s) select your predictor variables.
4. Make any other selections as required.
Example
The table below shows the results of a linear discriminant analysis predicting brand preference based on the attributes of the brand. The sub-title shows the predictive accuracy of the model, which in this case is extremely poor, at approximately 7%. The colored shading shows the differences between the means by group. It shows, for example, that the 1,799 Coca-Cola drinkers in the sample has significantly lower ratings of health-conscious, older, and traditional (these are the only significant differences, when compared to the mean, which is why they are in bold. We can also see that there are some significant differences relating to Pepsi. The R-Squared column shows the proportion of variance within each row that is explained by the groups; in all cases it is very poor. See Analysis of Variance - One-Way MANOVA for more detail on the interpretation of the table.
There are two reasons why this model is particularly poor:
- The relationship between the predictors and the outcome is weak.
- The Prior is at Equal, which assumes that the group sizes in the population are equal. In this example, Coca-Cola is by far the biggest group, so the prior causes the predicted accuracy to be poor.
Options
The inputs used to generate the Linear Discriminant Analysis are shown below.
Outcome The variable to be predicted by the predictor variables.
Predictors The numeric variable(s) to predict the outcome.
Algorithm The machine learning algorithm. Defaults to Linear Discriminant Analysis but may be changed to other machine learning methods.
Output
- Means Produces a table showing the means by category, and assorted statistics to evaluate the LDA.
- Detail More detailed diagnostics, from the lda function in the R MASS package.
- Prediction-Accuracy Table Produces a table relating the observed and predicted outcome. Also known as a confusion matrix.
- Scatterplot A two-dimensional scatterplot of the group centroids in the space of the first two discriminant function variables. This shows which groups are separated by the first two discriminant function variables. Also plotted are the correlations between the predictor variables and the first two discriminant function variables. The group centroids are scaled to appear on the same scale as the correlations.
- Moonplot A two-dimensional moonplot, using the same assumptions as the scatterplot.
Outcome color Color of group centroids in Scatterplot output.
Predictors color Color of variable correlations in Scatterplot output.
Missing data See Missing Data Options.
Variable names Displays Variable Names in the output instead of labels.
Prior The prior probabilities used in computing the probabilities of group membership of the Outcome (Machine Learning - Save Variable(s) - Probabilities of Each Response). Note that in the main R package for discriminant analysis (MASS:lda), the priors are also used in fitting the model, and this means that results differ between the normal R discriminant analysis and the results in this procedure. This procedure matches the results from SPSS.
- Equal The prior probabilities are assumed to be equal for each group of the Outcome.
- Observed Prior computed based on the current (weighted) group sizes. This is the default.
Random seed Seed used to initialize the (pseudo)random number generator for the model fitting algorithm. Different seeds may lead to slightly different answers, but should normally not make a large difference.
Increase allowed output size Check this box if you encounter a warning message "The R output had size XXX MB, exceeding the 128 MB limit..." and you need to reference the output elsewhere in your document; e.g., to save predicted values to a Data Set or examine diagnostics.
Maximum allowed size for output (MB). This control only appears if Increase allowed output size is checked. Use it to set the maximum allowed size for the regression output in MegaBytes. The warning referred to above about the R output size will state the minimum size you need to increase to to return the full output. Note that having very many large outputs in one document or page may slow down the performance of your document and increase load times.
Weight. Where a weight has been set for the R Output, it will automatically applied when the model is estimated. By default, the weight is assumed to be a sampling weight, and the standard errors are estimated using Taylor series linearization (by contrast, in the Legacy Regression, weight calibration is used). See Weights, Effective Sample Size and Design Effects.
Filter The data is automatically filtered using any filters prior to estimating the model.
Additional options are available by editing the code.
DIAGNOSTICS
Coefficients of discriminant functions Creates a table of coefficients of linear discriminant functions for each class.
Prediction-Accuracy Table Creates a table showing the observed and predicted values, as a heatmap.
SAVE VARIABLE(S)
Discriminant Variables Creates a new question containing the discriminant variables.
Predicted Values Creates a new variable containing predicted values for each case in the data.
Probabilities of Each Response Creates new variables containing predicted probabilities of each response.
Acknowledgements
The algorithm used for fitting the LDA is a modification of MASS:lda, generalized to accommodate weights. The multcomp package is used to test comparisons (see also Regression - Generalized Linear Model, which describes the models that are used by multcomp). The survey package is used to compute the p for each of the variables in Means; a Wald test is used (regTermTest)
More information
See this post for a description of LDA.
See this post for a practical guide of how to run LDA in Displayr.
Code
var controls = [];
// ALGORITHM
var algorithm = form.comboBox({ label: "Algorithm",
alternatives: ["CART", "Deep Learning", "Gradient Boosting", "Linear Discriminant Analysis",
"Random Forest", "Regression", "Support Vector Machine"],
name: "formAlgorithm", default_value: "Linear Discriminant Analysis",
prompt: "Machine learning or regression algorithm for fitting the model" });
controls.push(algorithm);
algorithm = algorithm.getValue();
var regressionType = "";
if (algorithm == "Regression") {
regressionTypeControl = form.comboBox({ label: "Regression type",
alternatives: ["Linear", "Binary Logit", "Ordered Logit", "Multinomial Logit", "Poisson",
"Quasi-Poisson", "NBD"],
name: "formRegressionType", default_value: "Linear",
prompt: "Select type according to outcome variable type" });
regressionType = regressionTypeControl.getValue();
controls.push(regressionTypeControl);
}
// DEFAULT CONTROLS
missing_data_options = ["Error if missing data", "Exclude cases with missing data", "Imputation (replace missing values with estimates)"];
// AMEND DEFAULT CONTROLS PER ALGORITHM
if (algorithm == "Support Vector Machine")
output_options = ["Accuracy", "Prediction-Accuracy Table", "Detail"];
if (algorithm == "Gradient Boosting")
output_options = ["Accuracy", "Importance", "Prediction-Accuracy Table", "Detail"];
if (algorithm == "Random Forest")
output_options = ["Importance", "Prediction-Accuracy Table", "Detail"];
if (algorithm == "Deep Learning")
output_options = ["Accuracy", "Prediction-Accuracy Table", "Cross Validation", "Network Layers"];
if (algorithm == "Linear Discriminant Analysis")
output_options = ["Means", "Detail", "Prediction-Accuracy Table", "Scatterplot", "Moonplot"];
if (algorithm == "CART") {
output_options = ["Sankey", "Tree", "Text", "Prediction-Accuracy Table", "Cross Validation"];
missing_data_options = ["Error if missing data", "Exclude cases with missing data",
"Use partial data", "Imputation (replace missing values with estimates)"];
}
if (algorithm == "Regression") {
if (regressionType == "Multinomial Logit")
output_options = ["Summary", "Detail", "ANOVA"];
else if (regressionType == "Linear")
output_options = ["Summary", "Detail", "ANOVA", "Relative Importance Analysis", "Shapley Regression", "Jaccard Coefficient", "Correlation", "Effects Plot"];
else
output_options = ["Summary", "Detail", "ANOVA", "Relative Importance Analysis", "Effects Plot"];
}
// COMMON CONTROLS FOR ALL ALGORITHMS
var outputControl = form.comboBox({ label: "Output", prompt: "The type of output used to show the results",
alternatives: output_options, name: "formOutput",
default_value: output_options[0] });
controls.push(outputControl);
var output = outputControl.getValue();
if (algorithm == "Regression") {
if (regressionType == "Linear") {
if (output == "Jaccard Coefficient" || output == "Correlation")
missing_data_options = ["Error if missing data", "Exclude cases with missing data", "Use partial data (pairwise correlations)"];
else
missing_data_options = ["Error if missing data", "Exclude cases with missing data", "Dummy variable adjustment", "Use partial data (pairwise correlations)", "Multiple imputation"];
}
else
missing_data_options = ["Error if missing data", "Exclude cases with missing data", "Dummy variable adjustment", "Multiple imputation"];
}
var missingControl = form.comboBox({ label: "Missing data",
alternatives: missing_data_options, name: "formMissing", default_value: "Exclude cases with missing data",
prompt: "Options for handling cases with missing data" });
var missing = missingControl.getValue();
controls.push(missingControl);
controls.push(form.checkBox({ label: "Variable names", name: "formNames", default_value: false, prompt: "Display names instead of labels" }));
// CONTROLS FOR SPECIFIC ALGORITHMS
if (algorithm == "Support Vector Machine")
controls.push(form.textBox({ label: "Cost", name: "formCost", default_value: 1, type: "number",
prompt: "High cost produces a complex model with risk of overfitting, low cost produces a simpler mode with risk of underfitting" }));
if (algorithm == "Gradient Boosting") {
controls.push(form.comboBox({ label: "Booster",
alternatives: ["gbtree", "gblinear"], name: "formBooster", default_value: "gbtree",
prompt: "Boost tree or linear underlying models" }));
controls.push(form.checkBox({ label: "Grid search", name: "formSearch", default_value: false,
prompt: "Search for optimal hyperparameters" }));
}
if (algorithm == "Random Forest")
if (output == "Importance")
controls.push(form.checkBox({ label: "Sort by importance", name: "formImportance", default_value: true }));
if (algorithm == "Deep Learning") {
controls.push(form.numericUpDown({ name: "formEpochs", label: "Maximum epochs", default_value: 10, minimum: 1, maximum: Number.MAX_SAFE_INTEGER,
prompt: "Number of rounds of training" }));
controls.push(form.textBox({ name: "formHiddenLayers", label: "Hidden layers", prompt: "Comma delimited list of the number of nodes in each hidden layer", required: true }));
controls.push(form.checkBox({ label: "Normalize predictors", name: "formNormalize", default_value: true,
prompt: "Normalize to zero mean and unit variance" }));
}
if (algorithm == "Linear Discriminant Analysis") {
if (output == "Scatterplot") {
controls.push(form.colorPicker({ label: "Outcome color", name: "formOutColor", default_value: "#5B9BD5" }));
controls.push(form.colorPicker({ label: "Predictors color", name: "formPredColor", default_value: "#ED7D31" }));
}
controls.push(form.comboBox({ label: "Prior", alternatives: ["Equal", "Observed",], name: "formPrior", default_value: "Observed",
prompt: "Probabilities of group membership" }));
}
if (algorithm == "CART") {
controls.push(form.comboBox({ label: "Pruning", alternatives: ["Minimum error", "Smallest tree", "None"],
name: "formPruning", default_value: "Minimum error",
prompt: "Remove nodes after tree has been built" }));
controls.push(form.checkBox({ label: "Early stopping", name: "formStopping", default_value: false,
prompt: "Stop building tree when fit does not improve" }));
controls.push(form.comboBox({ label: "Predictor category labels", alternatives: ["Full labels", "Abbreviated labels", "Letters"],
name: "formPredictorCategoryLabels", default_value: "Abbreviated labels",
prompt: "Labelling of predictor categories in the tree" }));
controls.push(form.comboBox({ label: "Outcome category labels", alternatives: ["Full labels", "Abbreviated labels", "Letters"],
name: "formOutcomeCategoryLabels", default_value: "Full labels",
prompt: "Labelling of outcome categories in the tree" }));
controls.push(form.checkBox({ label: "Allow long-running calculations", name: "formLongRunningCalculations", default_value: false,
prompt: "Allow predictors with more than 30 categories" }));
}
var stacked_check = false;
if (algorithm == "Regression") {
if (missing == "Multiple imputation")
controls.push(form.dropBox({ label: "Auxiliary variables",
types: ["Variable: Numeric, Date, Money, Categorical, OrderedCategorical"],
name: "formAuxiliaryVariables", required: false, multi: true,
prompt: "Additional variables to use when imputing missing values" }));
controls.push(form.comboBox({ label: "Correction", alternatives: ["None", "False Discovery Rate", "Bonferroni"], name: "formCorrection",
default_value: "None", prompt: "Multiple comparisons correction applied when computing p-values of post-hoc comparisons" }));
var is_RIA_or_shapley = output == "Relative Importance Analysis" || output == "Shapley Regression";
var is_Jaccard_or_Correlation = output == "Jaccard Coefficient" || output == "Correlation";
if (regressionType == "Linear" && missing != "Use partial data (pairwise correlations)" && missing != "Multiple imputation")
controls.push(form.checkBox({ label: "Robust standard errors", name: "formRobustSE", default_value: false,
prompt: "Standard errors are robust to violations of assumption of constant variance" }));
if (is_RIA_or_shapley)
controls.push(form.checkBox({ label: "Absolute importance scores", name: "formAbsoluteImportance", default_value: false,
prompt: "Show absolute instead of signed importances" }));
if (regressionType != "Multinomial Logit" && (is_RIA_or_shapley || is_Jaccard_or_Correlation || output == "Summary"))
controls.push(form.dropBox({ label: "Crosstab interaction", name: "formInteraction", types: ["Variable: Numeric, Date, Money, Categorical, OrderedCategorical"],
required: false, prompt: "Categorical variable to test for interaction with other variables" }));
if (regressionType !== "Multinomial Logit")
controls.push(form.numericUpDown({ name: "formOutlierProportion", label: "Automated outlier removal percentage", default_value: 0,
minimum: 0, maximum: 49.9, increment: 0.1,
prompt: "Data points removed and model refitted based on the residual values in the model using the full dataset" }));
stacked_check_box = form.checkBox({ label: "Stack data", name: "formStackedData", default_value: false,
prompt: "Allow input into the Outcome control to be a single multi variable and Predictors to be a single grid variable" });
stacked_check = stacked_check_box.getValue();
controls.push(stacked_check_box);
}
controls.push(form.numericUpDown({ name: "formSeed", label: "Random seed", default_value: 12321, minimum: 1, maximum: Number.MAX_SAFE_INTEGER,
prompt: "Initializes randomization for imputation and certain algorithms" }));
let allowLargeOutputsCtrl = form.checkBox({ label: "Increase allowed output size",
name: "formAllowLargeOutputs", default_value: false,
prompt: "Increase the limit on the maximum size allowed for the output to fix warnings about it being too large" });
controls.push(allowLargeOutputsCtrl);
if (allowLargeOutputsCtrl.getValue())
controls.push(form.numericUpDown({ name: "formMaxOutputSize", label: "Maximum allowed size for output (MB)", default_value: 128, minimum: 1, maximum: Number.MAX_SAFE_INTEGER,
prompt: "The maximum allowed size for the returned output in MB. Very large outputs may impact document performance" }));
var outcome = form.dropBox({ label: "Outcome",
types: [stacked_check ? "VariableSet: BinaryMulti, NominalMulti, OrdinalMulti, NumericMulti" : "Variable: Numeric, Date, Money, Categorical, OrderedCategorical"],
multi: false,
name: "formOutcomeVariable",
prompt: "Independent target variable to be predicted" });
var predictors = form.dropBox({ label: "Predictor(s)",
types: [stacked_check ? "VariableSet: BinaryGrid, NumericGrid" : "Variable: Numeric, Date, Money, Categorical, OrderedCategorical"],
name: "formPredictorVariables", multi: stacked_check ? false : true,
prompt: "Dependent input variables" });
controls.unshift(predictors);
controls.unshift(outcome);
form.setInputControls(controls);
var heading_text = "";
if (regressionType == "") {
heading_text = algorithm;
}
else {
heading_text = regressionType + " " + algorithm;
}
if (!!form.setObjectInspectorTitle)
form.setObjectInspectorTitle(heading_text, heading_text);
else
form.setHeading(heading_text);
library(flipMultivariates)
if (get0("formAllowLargeOutputs", ifnotfound = FALSE))
QAllowLargeResultObject(1e6*get0("formMaxOutputSize"))
WarnIfVariablesSelectedFromMultipleDataSets()
model <- MachineLearning(formula = if (isTRUE(get0("formStackedData"))) as.formula(NULL) else QFormula(formOutcomeVariable ~ formPredictorVariables),
algorithm = formAlgorithm,
weights = QPopulationWeight, subset = QFilter,
missing = formMissing,
output = if (formOutput == "Shapley Regression") "Shapley regression" else formOutput,
show.labels = !formNames,
seed = get0("formSeed"),
cost = get0("formCost"),
booster = get0("formBooster"),
grid.search = get0("formSearch"),
sort.by.importance = get0("formImportance"),
hidden.nodes = get0("formHiddenLayers"),
max.epochs = get0("formEpochs"),
normalize = get0("formNormalize"),
outcome.color = get0("formOutColor"),
predictors.color = get0("formPredColor"),
prior = get0("formPrior"),
prune = get0("formPruning"),
early.stopping = get0("formStopping"),
predictor.level.treatment = get0("formPredictorCategoryLabels"),
outcome.level.treatment = get0("formOutcomeCategoryLabels"),
long.running.calculations = get0("formLongRunningCalculations"),
type = get0("formRegressionType"),
auxiliary.data = get0("formAuxiliaryVariables"),
correction = get0("formCorrection"),
robust.se = get0("formRobustSE", ifnotfound = FALSE),
importance.absolute = get0("formAbsoluteImportance"),
interaction = get0("formInteraction"),
outlier.prop.to.remove = if (get0("formRegressionType", ifnotfound = "") != "Multinomial Logit") get0("formOutlierProportion")/100 else NULL,
stacked.data.check = get0("formStackedData"),
unstacked.data = if (isTRUE(get0("formStackedData"))) list(Y = get0("formOutcomeVariable"), X = get0("formPredictorVariables")) else NULL,
use.combined.scatter = TRUE)